SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets, BMC Research Notes
4.6 (239) · $ 21.50 · In stock


PDF) Whole-Genome Sequencing Confirms that Burkholderia pseudomallei Multilocus Sequence Types Common to Both Cambodia and Australia Are Due to Homoplasy

Dynamics of genetic variation in Transcription Factors and its implications for the evolution of regulatory networks in Bacteria

Generalizable characteristics of false-positive bacterial variant calls

Unprecedented Melioidosis Cases in Northern Australia Caused by an Asian Burkholderia pseudomallei Strain Identified by Using Large-Scale Comparative Genomics. - Abstract - Europe PMC

Using whole-genome sequencing and a pentaplex real-time PCR to characterize third-generation cephalosporin-resistant Enterobacteriaceae from Southeast Queensland, Australia
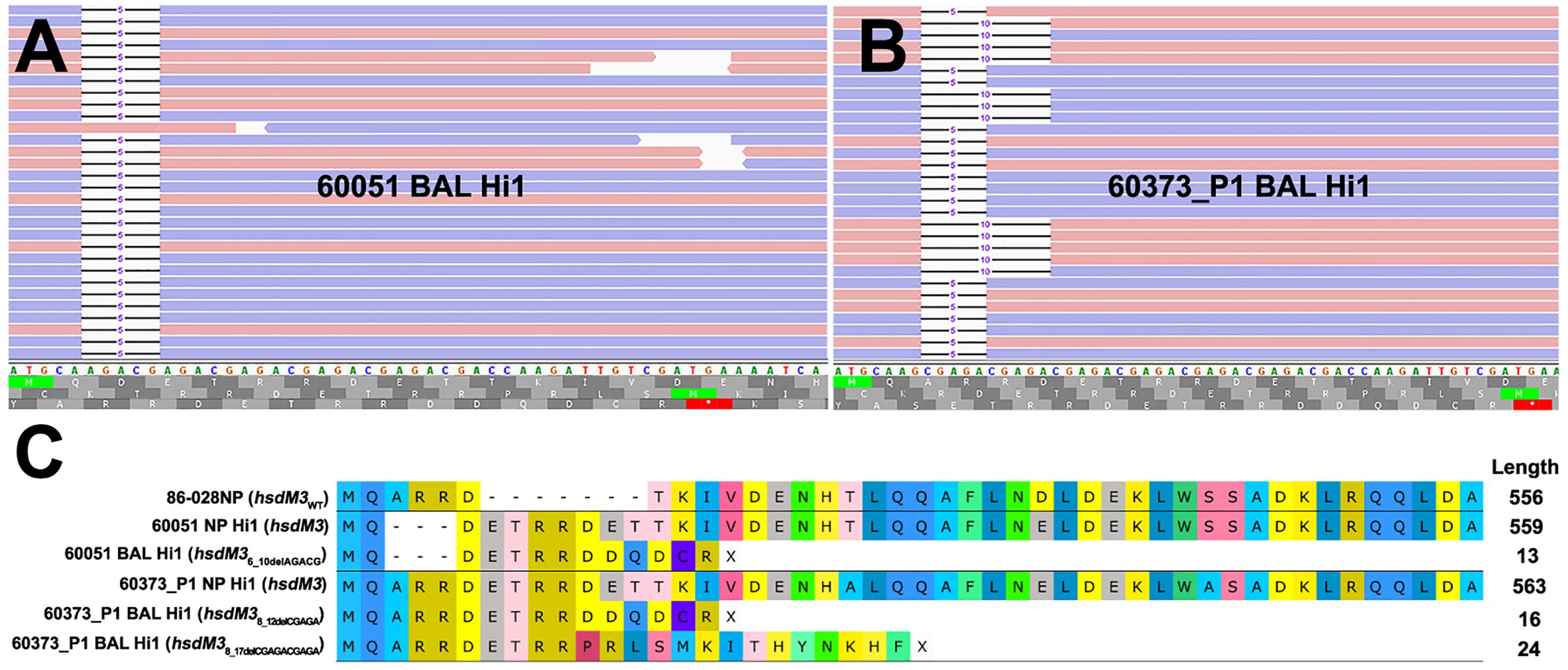
Frontiers Molecular Signatures of Non-typeable Haemophilus influenzae Lung Adaptation in Pediatric Chronic Lung Disease
Tracing the environmental footprint of the Burkholderia pseudomallei lipopolysaccharide genotypes in the tropical “Top End” of the Northern Territory, Australia

Comparative genomics confirms a rare melioidosis human-to-human transmission event and reveals incorrect phylogenomic reconstruction due to polyclonality

Whole genome sequencing and comparative genomic analysis of oleaginous red yeast Sporobolomyces pararoseus NGR identifies candidate genes for biotechnological potential and ballistospores-shooting, BMC Genomics
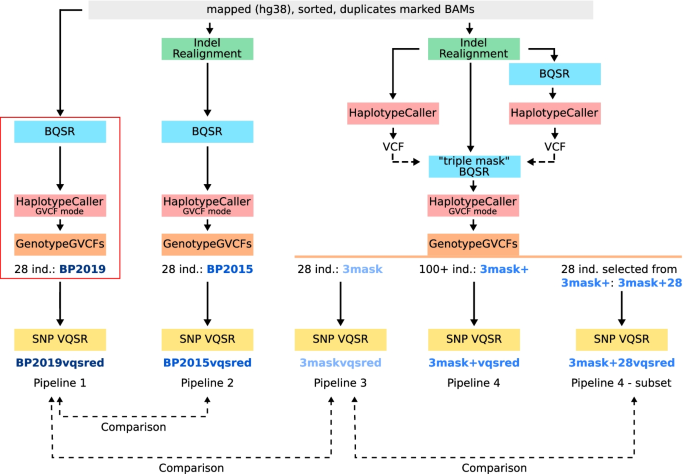
Comparison of sequencing data processing pipelines and application to underrepresented African human populations, BMC Bioinformatics
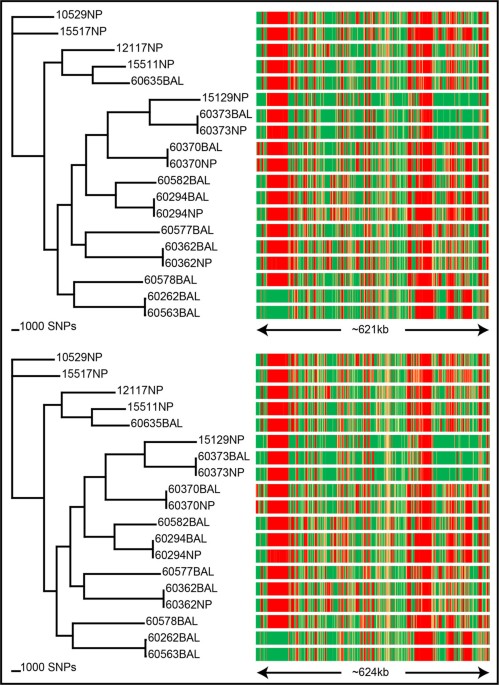
SPANDx: a genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets, BMC Research Notes







